Navigation
What is iHypoxia
The iHypoxia is a hypoxia associated protein database, which collated 865 literature for 5 mammals hypoxia research. In total, 458,481 quantification events on 39,685 hypoxia-regulated proteins under more than 390 conditions/states and ~10,000 candidate genes were collected and integrated in the database. Then, the data were expanded to 432,298 proteins in 48 animals by orthologous search. Various browse and search options are provided in the database, and the detailed information is organized and visualized for queries.
Hypoxia is a condition or state in which insufficient levels of oxygen are supplied to cells or tissues. The research of hypoxia mainly studies physiological and pathological changes under hypoxic conditions and reveal its role in diseases, tumors, biological processes and so on.
How to use Hypoxia
Quick Search
In the home page of iHypoxia, users can start a quick search with the keyword(s) of interest or simply click an example that we provided above the input field and search button to perform an example search.
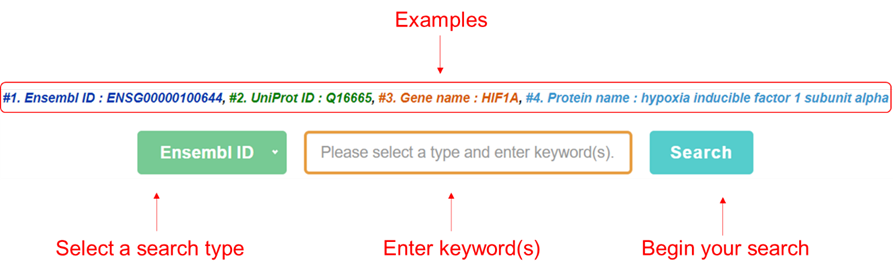

Browse
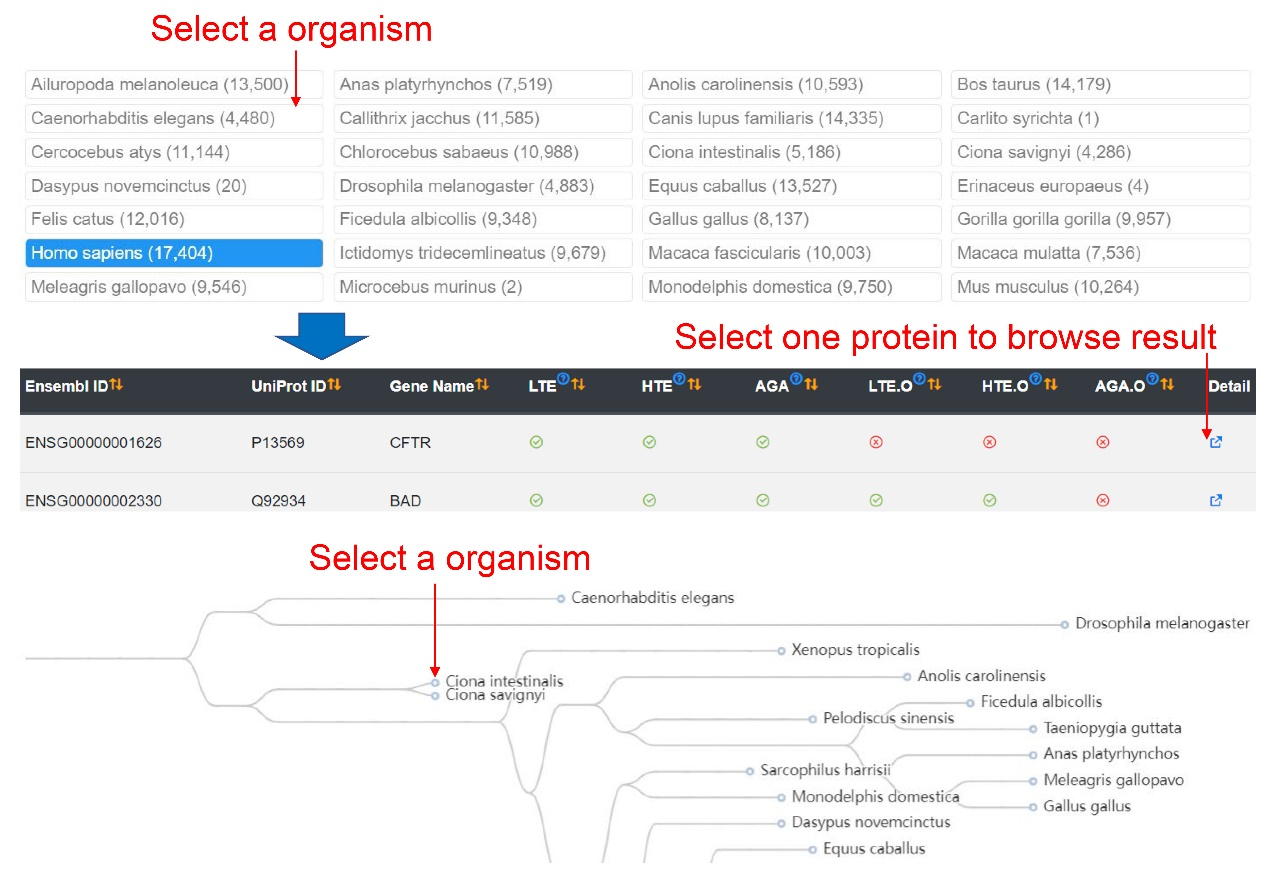
On the browse page, all animal species in iHypoxia are presented in alphabetical order. To view the proteins in species of interest, users can click the corresponding species, and a table with the basic information of the Ensembl ID, UniProt ID, gene name, and experiment types will be shown in the list below. Users can click the ‘Detail’ link to view detailed information.
Additionally, an evolutionary tree showing the phylogenetic relationships of these 48 animals is diagrammed and users can directly click the animal of interest to view corresponding proteins.
Search
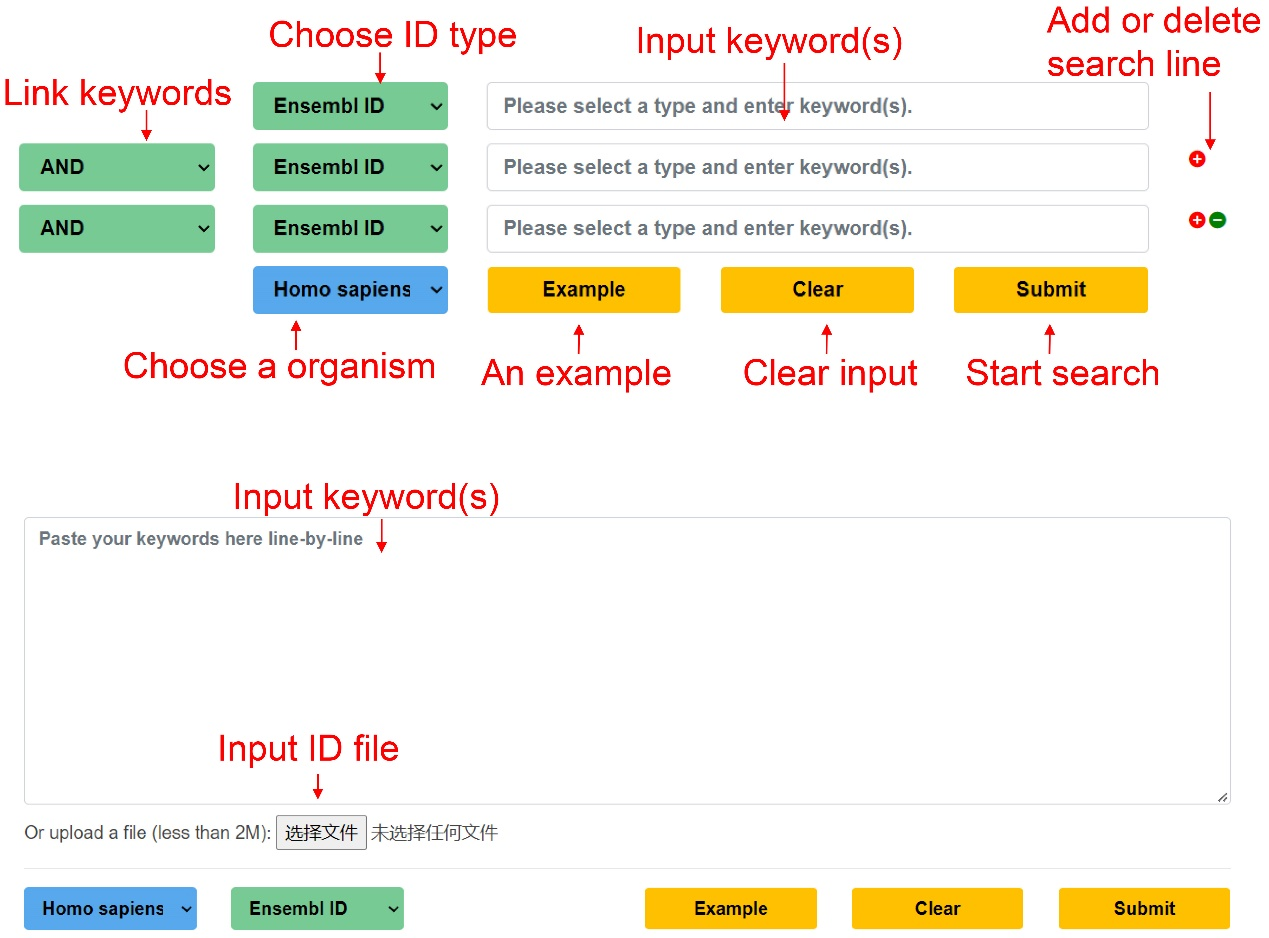
On the search page, we provided two search approaches including advanced search and batch search. For the option ‘Advanced search’, various keywords could be combined through the operators of "AND", "OR" and "NOT" to perform a query with different search keywords.
For the option ‘Batch search’, users can paste numerous keywords, such as Ensembl IDs and gene names in a line-by-line format, or upload a file for batch searching.
Result
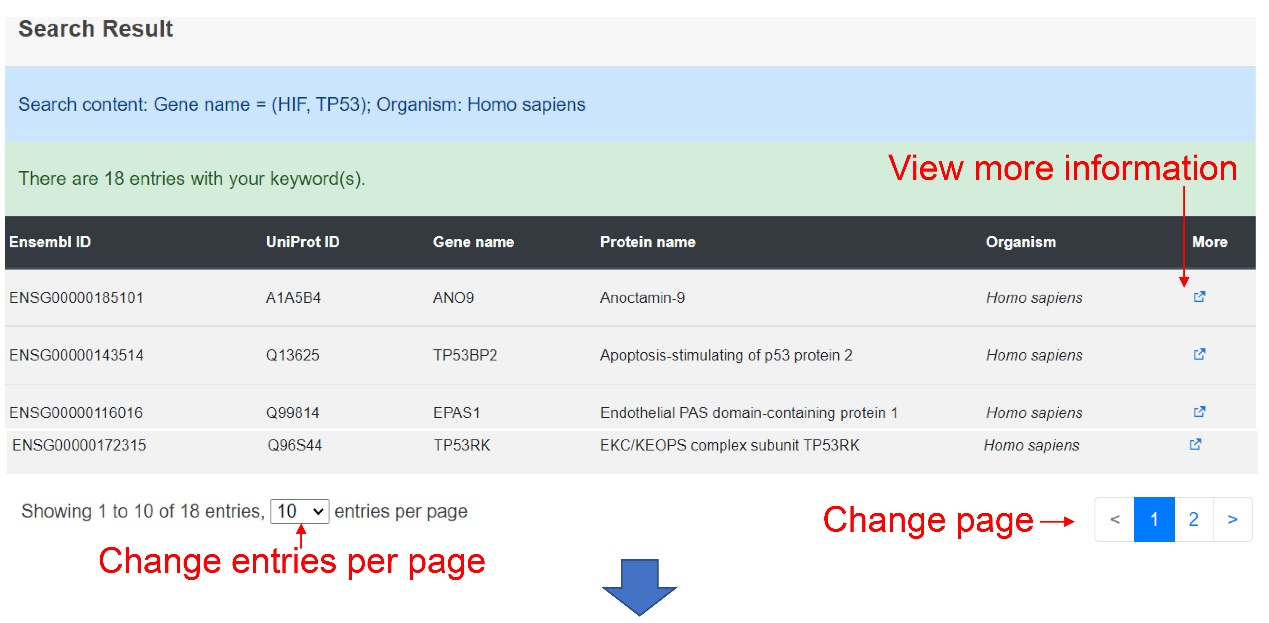
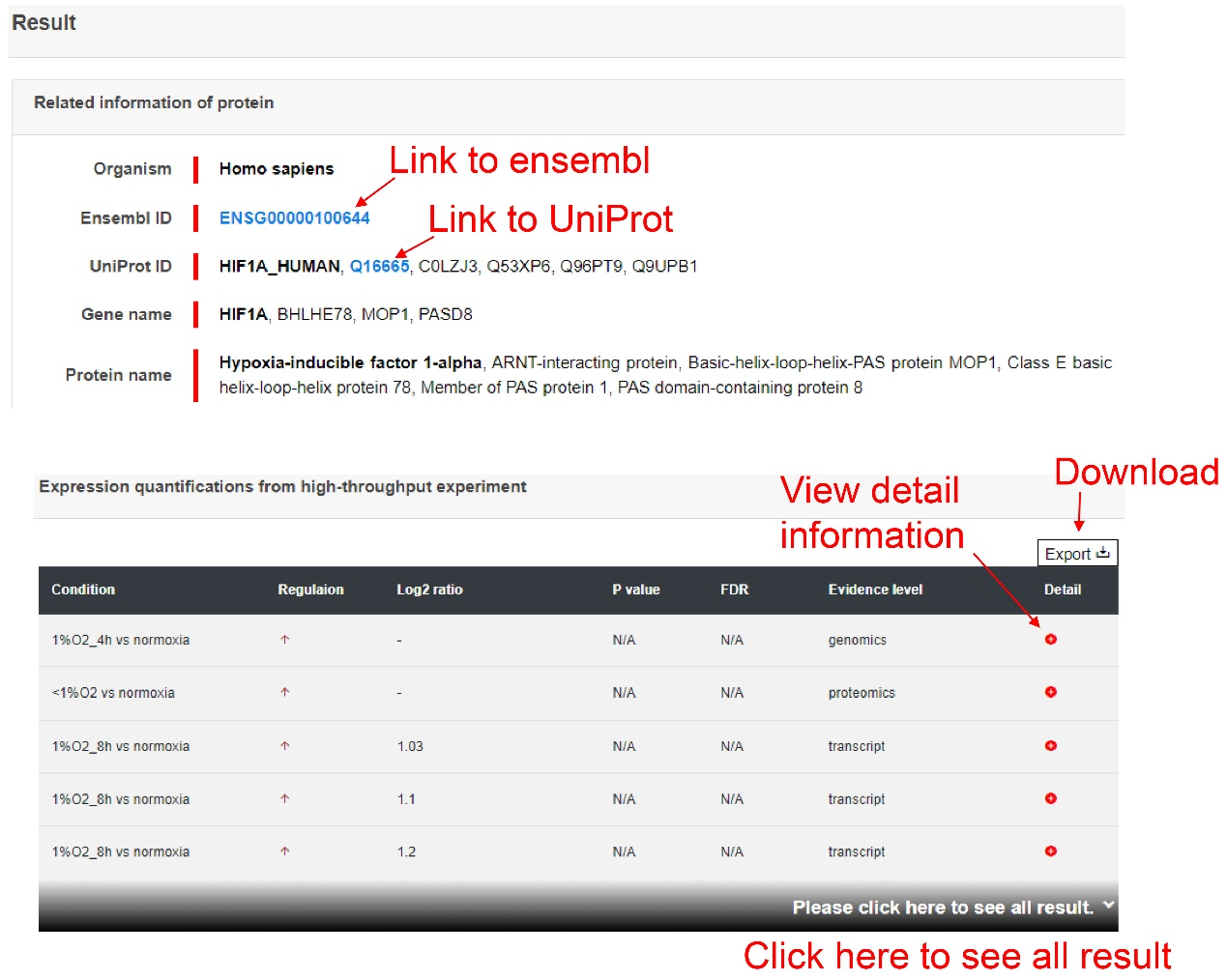
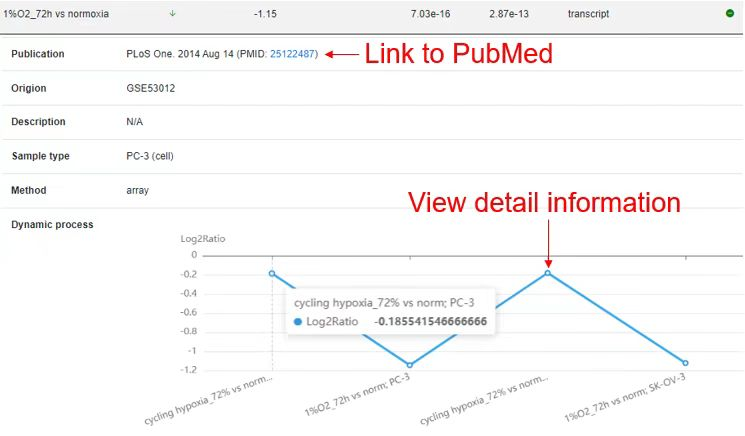
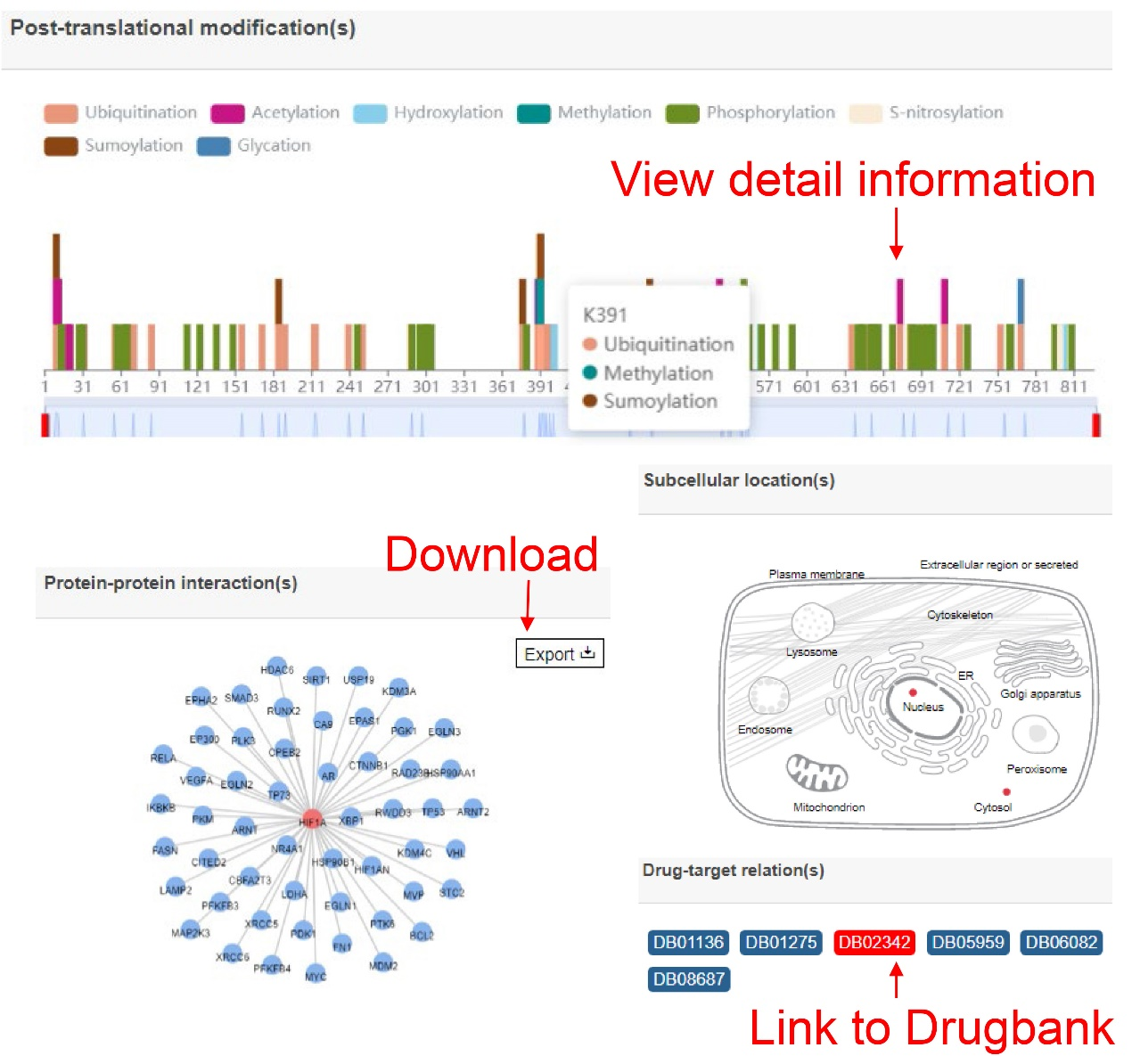
The result page shows the results that users searched and users can get more detailed results by click the 'More' button.


Hypoxia reference resources
Ensembl
Ensembl is a genome browser for vertebrate genomes that supports researches in comparative genomics, evolution, sequence variation and transcriptional regulation. Ensembl annotates genes, computes multiple alignments, predicts regulatory function and collects disease data. Ensembl tools include BLAST, BLAT, BioMart and the Variant Effect Predictor (VEP) for all supported species.
http://www.ensembl.org/UniProt
The mission of UniProt is to provide the scientific community with a comprehensive, high-quality and freely accessible resource of protein sequence and functional information.
https://www.uniprot.org/HypoxiaDB
HypoxiaDB is a first ever manually-curated non-redundant catalogue of human hypoxia-regulated proteins with a goal of collecting proteins whose expression patterns are altered in hypoxic conditions. It is a highly curated database providing information on expression pattern changes under different levels and duration of hypoxia. HypoxiaDB currently contains more than 72,000 entries taken on 3,500 differentially regulated proteins extracted from 73 peer-reviewed publications and ~1,500 supporting publications.
http://www.hypoxiadb.com/HRGFish
HRGFish is a database of hypoxia responsive genes in fishes, and covers 818 gene sequences and 35 gene types from 38 fishes.
http://mail.nbfgr.res.in/HRGFishGene Ontology (GO)
The Gene Ontology (GO) knowledgebase is the world’s largest source of information on the functions of genes. This knowledge is both human-readable and machine-readable, and is a foundation for computational analysis of large-scale molecular biology and genetics experiments in biomedical research.
http://geneontology.org/